Using XGboost to Predict Dyspnea in COVID-19 Survivors
By Saulo Gil
January 6, 2024
Introduction
Dyspnea is a medical term that refers to the sensation of shortness of breath or difficulty breathing. It is a common symptom of heart and lung diseases, such as asthma, pneumonia, and chronic obstructive pulmonary disease (COPD).
In the context of COVID-19, dyspnea is one of the most common symptoms experienced by infected individuals, but it remains even after acute COVID-19 is solved. According to a study published in BMC Pulmonary Medicine, 45% of COVID-19 pneumonia survivors admitted to the hospital experience dyspnea even after 1 year from hospital discharge. In this sense, identifying patients at risk of presenting dyspnea is crucial for early management.
XGBoost (eXtreme Gradient Boosting) is a very popular ML algorithm. Its popularity is due to the fact that has recently been dominating applied machine learning and Kaggle competitions for structured or tabular data. Therefore, the implementation of ML algorithm XGBoost in clinical setting may be useful for predicting COVID-19 survivors at the risk of dyspnea following discharge.
Herein, we utilized the XGBoost to predict dyspnea in COVID-19 survivors 6-9 months after hospital discharge
Packages
# Pacotes
library(tidyverse)
library(tidymodels)
library(GGally)
library(DataExplorer)
library(vip)
library(doParallel)Reading dataset
The dataset consist in a real dataset collected in the largest tertiary hospital of Latin America (Clinical Hospital, School of Medicine of the University of Sao Paulo).
# Lendo a base
dispneia <- read.csv2('df_ajustada.csv')Spliting dataset - Train/Test
Since our dataset is size-limited, I opted to separate 80% of the sample size for training.
set.seed(32)
dispneia_initial_split <- initial_split(dispneia,
prop = 0.8,
strata = "dispneia")
dispneia_initial_split## <Training/Testing/Total>
## <592/149/741># train and test datasets
dispneia_train <- training(dispneia_initial_split)
dispneia_test <- testing(dispneia_initial_split)Exploratory analysis
Overview
skimr::skim(dispneia_train)| Name | dispneia_train |
| Number of rows | 592 |
| Number of columns | 47 |
| _______________________ | |
| Column type frequency: | |
| character | 42 |
| numeric | 5 |
| ________________________ | |
| Group variables | None |
Variable type: character
| skim_variable | n_missing | complete_rate | min | max | empty | n_unique | whitespace |
|---|---|---|---|---|---|---|---|
| idade | 0 | 1.00 | 5 | 9 | 0 | 2 | 0 |
| sexo | 0 | 1.00 | 4 | 6 | 0 | 2 | 0 |
| adm_uti | 0 | 1.00 | 2 | 3 | 0 | 2 | 0 |
| tempo_uti | 0 | 1.00 | 9 | 16 | 0 | 3 | 0 |
| vmi | 0 | 1.00 | 2 | 3 | 0 | 2 | 0 |
| tempo_vmi | 0 | 1.00 | 9 | 17 | 0 | 3 | 0 |
| dispneia | 0 | 1.00 | 2 | 3 | 0 | 2 | 0 |
| tosse | 13 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| fraqueza | 7 | 0.99 | 2 | 3 | 0 | 2 | 0 |
| dific_caminhar | 7 | 0.99 | 2 | 3 | 0 | 2 | 0 |
| angina | 21 | 0.96 | 2 | 3 | 0 | 2 | 0 |
| dor_articular | 12 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| dor_abdominal | 12 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| diarreia | 14 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| nausea | 9 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| perda_consciencia | 12 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| inapetencia | 7 | 0.99 | 2 | 3 | 0 | 2 | 0 |
| parestesia | 8 | 0.99 | 2 | 3 | 0 | 2 | 0 |
| obstrucao_nasal | 16 | 0.97 | 2 | 3 | 0 | 2 | 0 |
| perda_paladar | 25 | 0.96 | 2 | 3 | 0 | 2 | 0 |
| anosmia | 18 | 0.97 | 2 | 3 | 0 | 2 | 0 |
| fadiga | 0 | 1.00 | 2 | 3 | 0 | 2 | 0 |
| cefaleia | 7 | 0.99 | 2 | 3 | 0 | 2 | 0 |
| trail_making_a_persistent_fup_incident | 73 | 0.88 | 2 | 3 | 0 | 2 | 0 |
| perda_memoria | 47 | 0.92 | 2 | 3 | 0 | 2 | 0 |
| TEPT | 11 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| perda_auditiva | 11 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| ruido_ouvido | 10 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| vertigem | 9 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| edema | 14 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| nocturia | 15 | 0.97 | 2 | 3 | 0 | 2 | 0 |
| dor_corpo | 9 | 0.98 | 2 | 3 | 0 | 2 | 0 |
| insonia | 0 | 1.00 | 2 | 3 | 0 | 2 | 0 |
| depressao | 64 | 0.89 | 2 | 3 | 0 | 2 | 0 |
| ansiedade | 64 | 0.89 | 2 | 3 | 0 | 2 | 0 |
| tempo_internacao | 0 | 1.00 | 9 | 11 | 0 | 2 | 0 |
| tabagismo | 4 | 0.99 | 2 | 3 | 0 | 2 | 0 |
| obesidade | 2 | 1.00 | 2 | 3 | 0 | 2 | 0 |
| hipertensao | 0 | 1.00 | 2 | 3 | 0 | 2 | 0 |
| diabetes | 0 | 1.00 | 2 | 3 | 0 | 2 | 0 |
| gravidade_adm | 0 | 1.00 | 5 | 8 | 0 | 4 | 0 |
| nivel_atf | 3 | 0.99 | 5 | 7 | 0 | 2 | 0 |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| espiro_cvf_pred | 86 | 0.85 | 83.46 | 16.51 | 22 | 73.00 | 84 | 94.00 | 132 | ▁▂▇▆▁ |
| espiro_cvf_vef1_pred | 86 | 0.85 | 101.78 | 9.91 | 43 | 97.00 | 103 | 107.00 | 129 | ▁▁▂▇▂ |
| espiro_vef_pred | 86 | 0.85 | 84.90 | 17.61 | 21 | 75.00 | 87 | 96.00 | 140 | ▁▂▇▅▁ |
| espiro_pefr_pred | 87 | 0.85 | 74.68 | 22.88 | 9 | 60.00 | 76 | 91.00 | 133 | ▁▃▇▆▂ |
| espiro_fef_pred | 86 | 0.85 | 101.74 | 39.00 | 0 | 75.25 | 100 | 125.75 | 282 | ▂▇▆▁▁ |
Missing values
plot_missing(dispneia_train)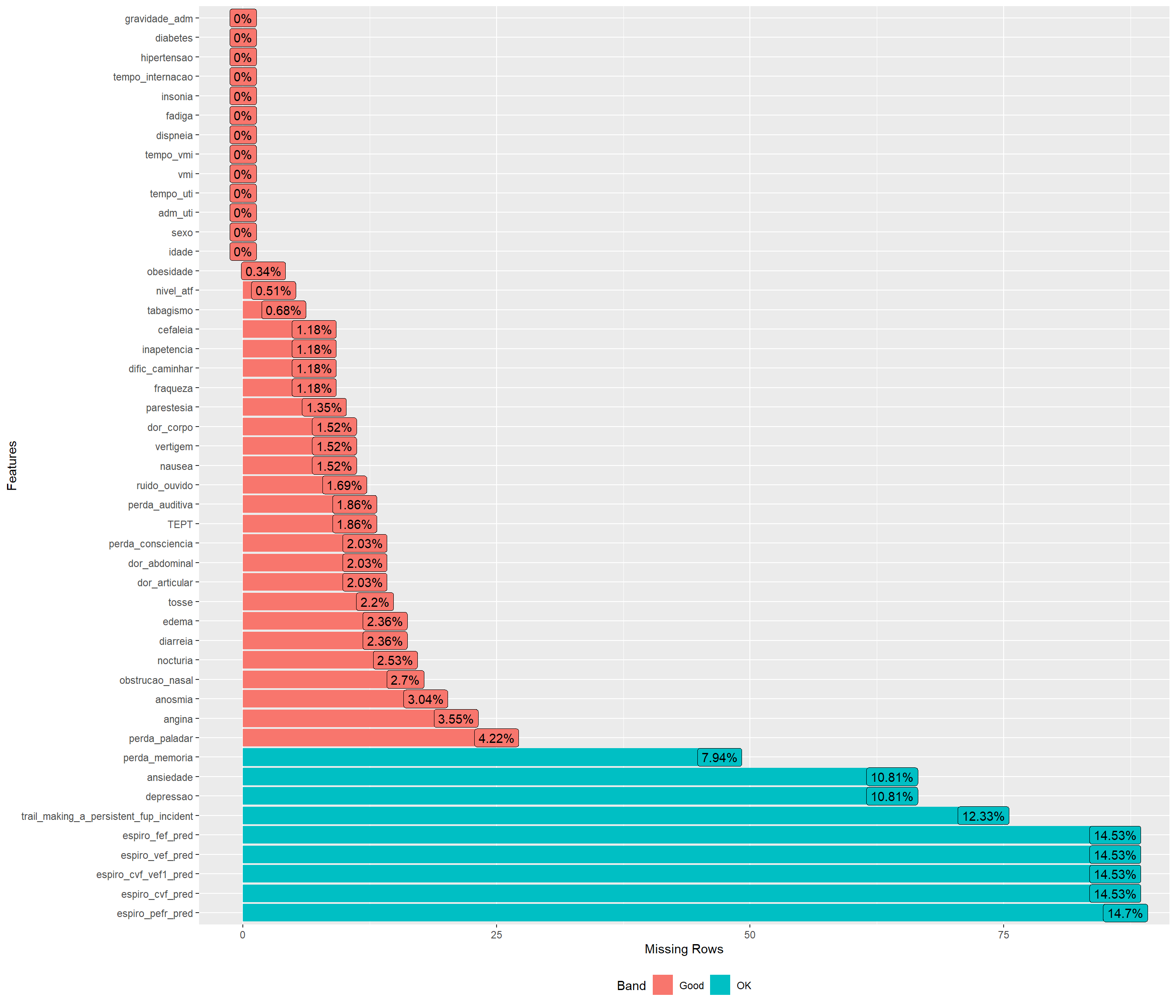
Checking the correlation and distribution of numerical features
dispneia_train |>
select(where(is.numeric)) |>
cor(use = "pairwise.complete.obs") |>
corrplot::corrplot(method = "number")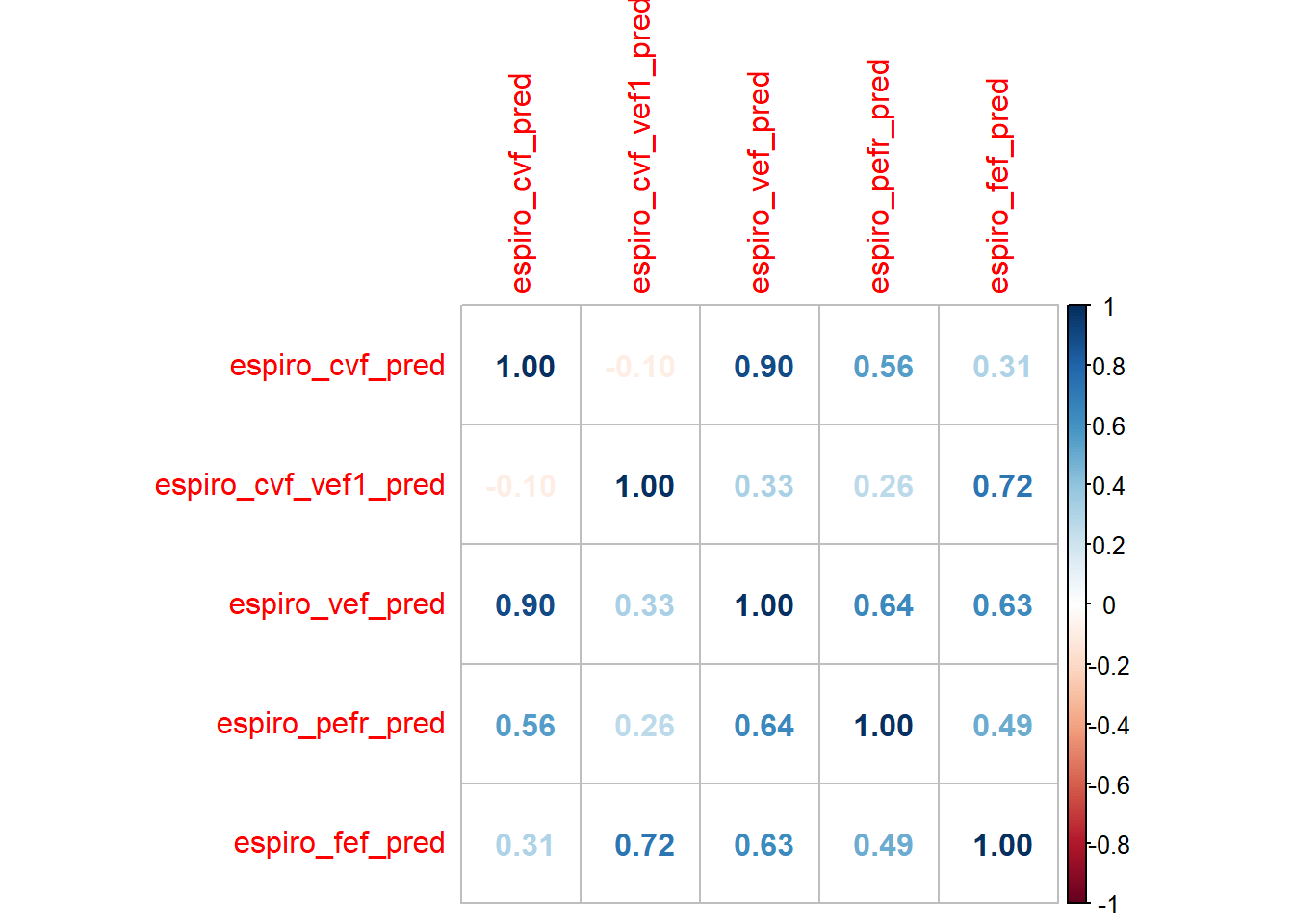
dispneia_train |>
select(where(is.numeric), dispneia) |>
ggpairs(aes(colour = dispneia))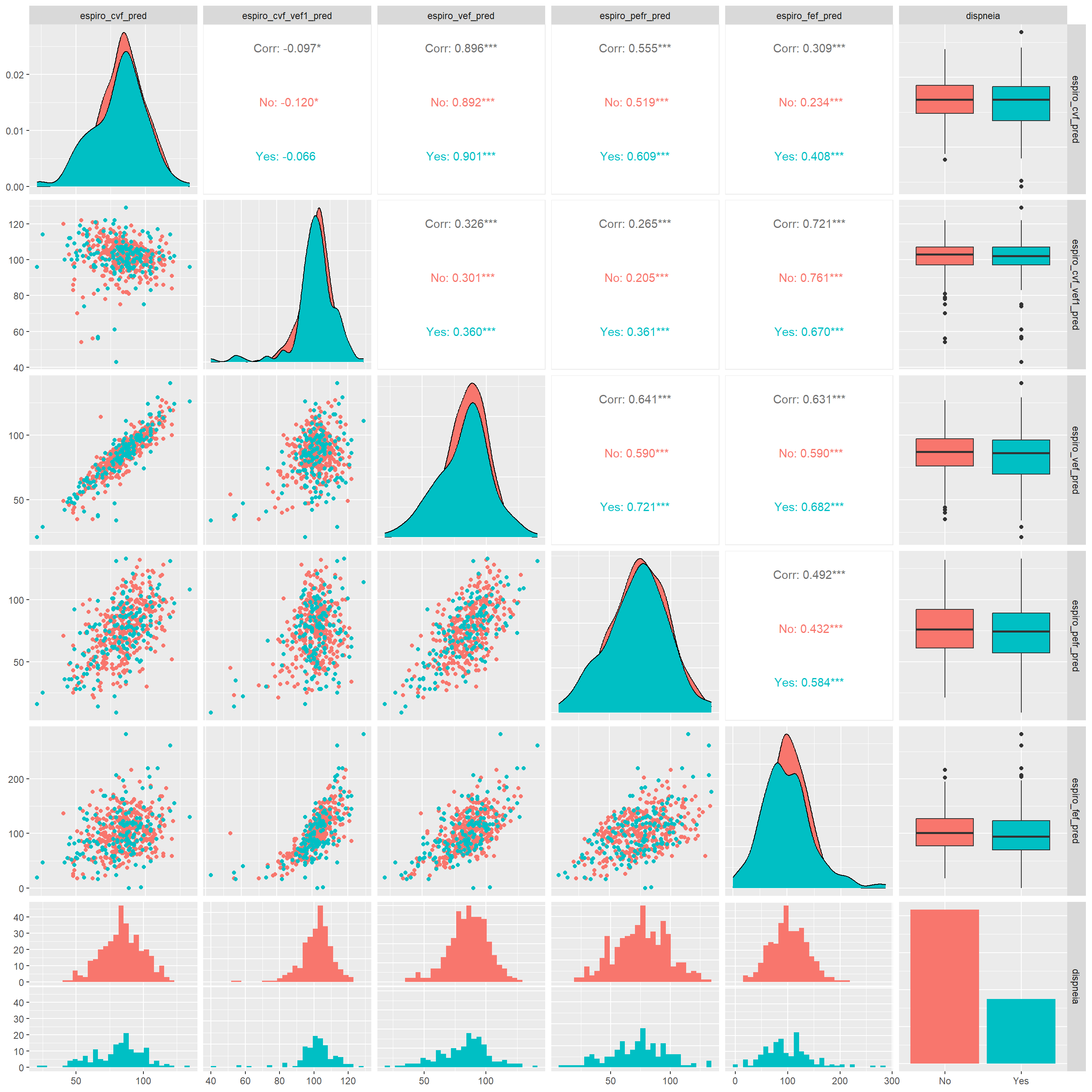
dispneia_train |>
select(c(where(is.numeric), dispneia)) |>
pivot_longer(-dispneia,
names_to = "Feature",
values_to = "Value") |>
ggplot(aes(y = dispneia,
x = Value,
fill = dispneia)) +
geom_boxplot() +
facet_wrap(~Feature,
scales = "free_x") +
ggtitle("Dyspnea vs. Numeric Features")## Warning: Removed 431 rows containing non-finite values (`stat_boxplot()`).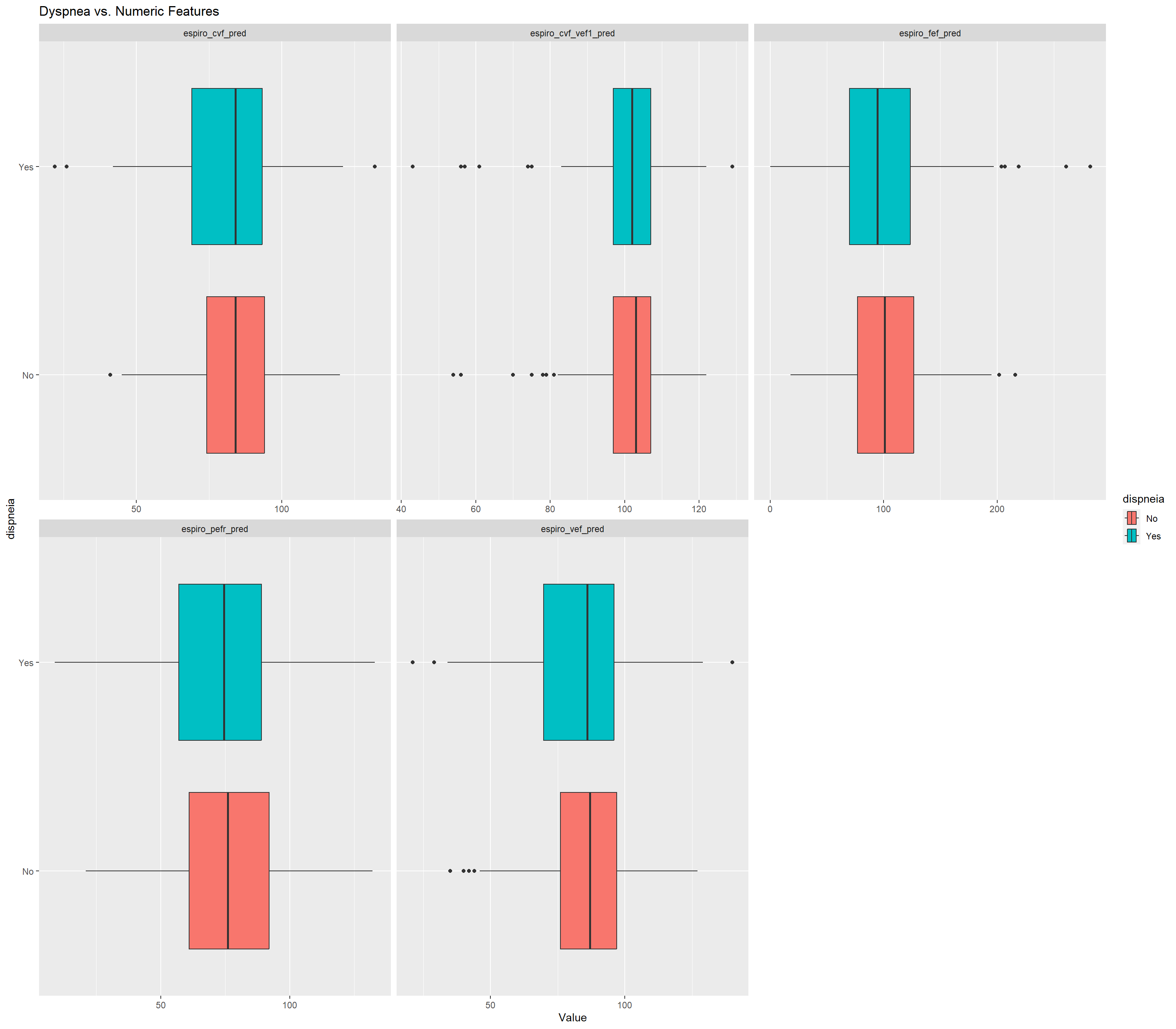
dispneia_train |>
select(c(where(is.numeric), dispneia)) |>
pivot_longer(-dispneia,
names_to = "Feature",
values_to = "Value") |>
ggplot(aes(x = Value,
colour = dispneia)) +
stat_ecdf() +
facet_wrap(~Feature, scales = "free_x") +
labs(title = "Dyspnea vs. Numeric Features",
subtitle = "Cumulative Distribution")## Warning: Removed 431 rows containing non-finite values (`stat_ecdf()`).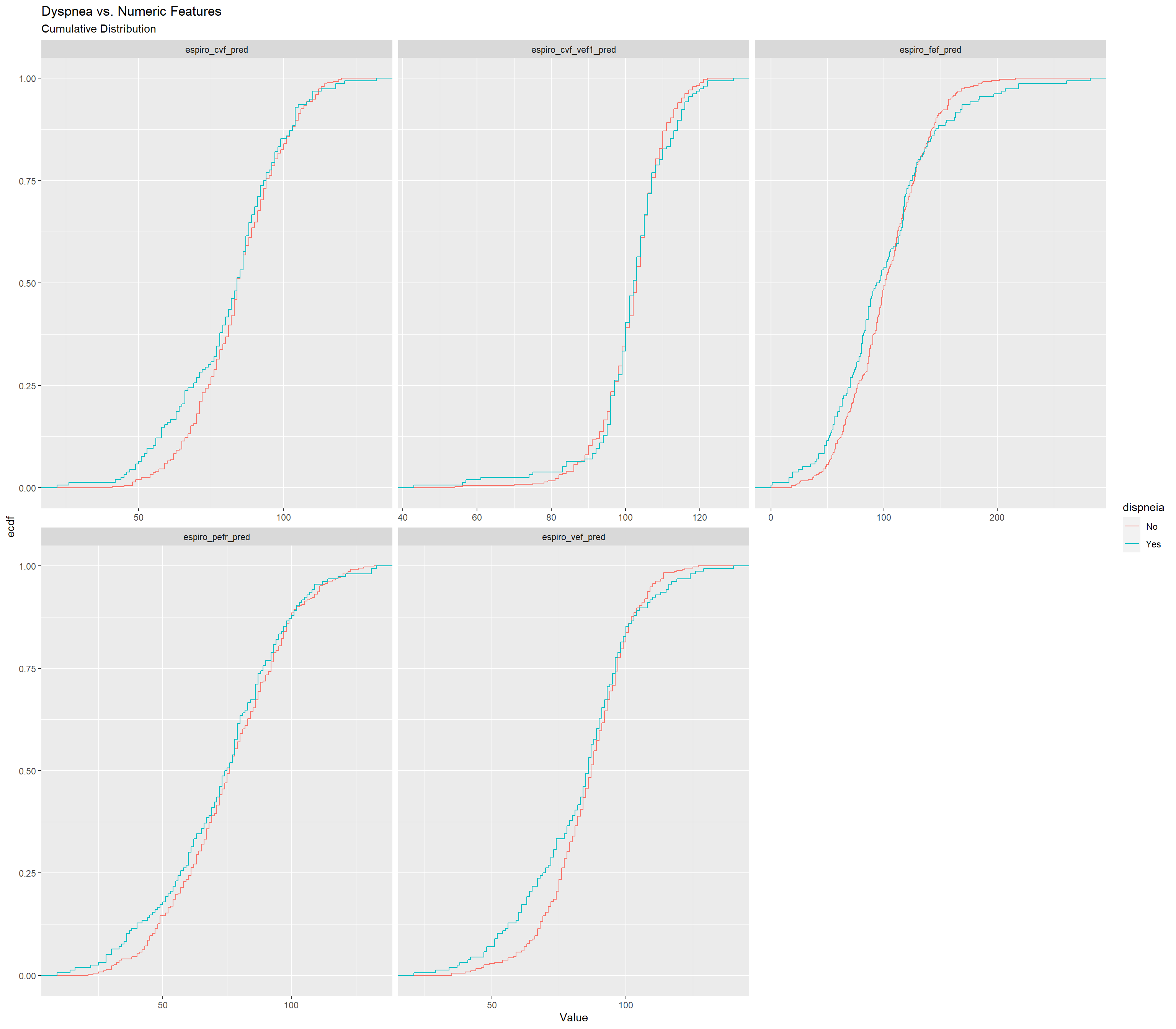
grafico_de_barras_das_vars_continuas <-
function(dados) {
dados |>
select(c(where(is.numeric), dispneia)) |>
pivot_longer(-dispneia,
names_to = "Feature",
values_to = "Value") |>
dplyr::group_by(Feature) |>
dplyr::mutate(
Value = factor(dplyr::ntile(Value, 10),
levels = 1:10)
) |>
dplyr::count(dispneia,
Feature,
Value) |>
ggplot(aes(y = (Value),
x = n,
fill = dispneia)) +
geom_col(position = "fill") +
geom_label(aes(label = n),
position = position_fill(vjust = 0.5)) +
facet_wrap(~Feature,
scales = "free_y",
ncol = 3) +
ggtitle("Dyspnea vs. Numeric Features")
}
grafico_de_barras_das_vars_continuas(dispneia_train)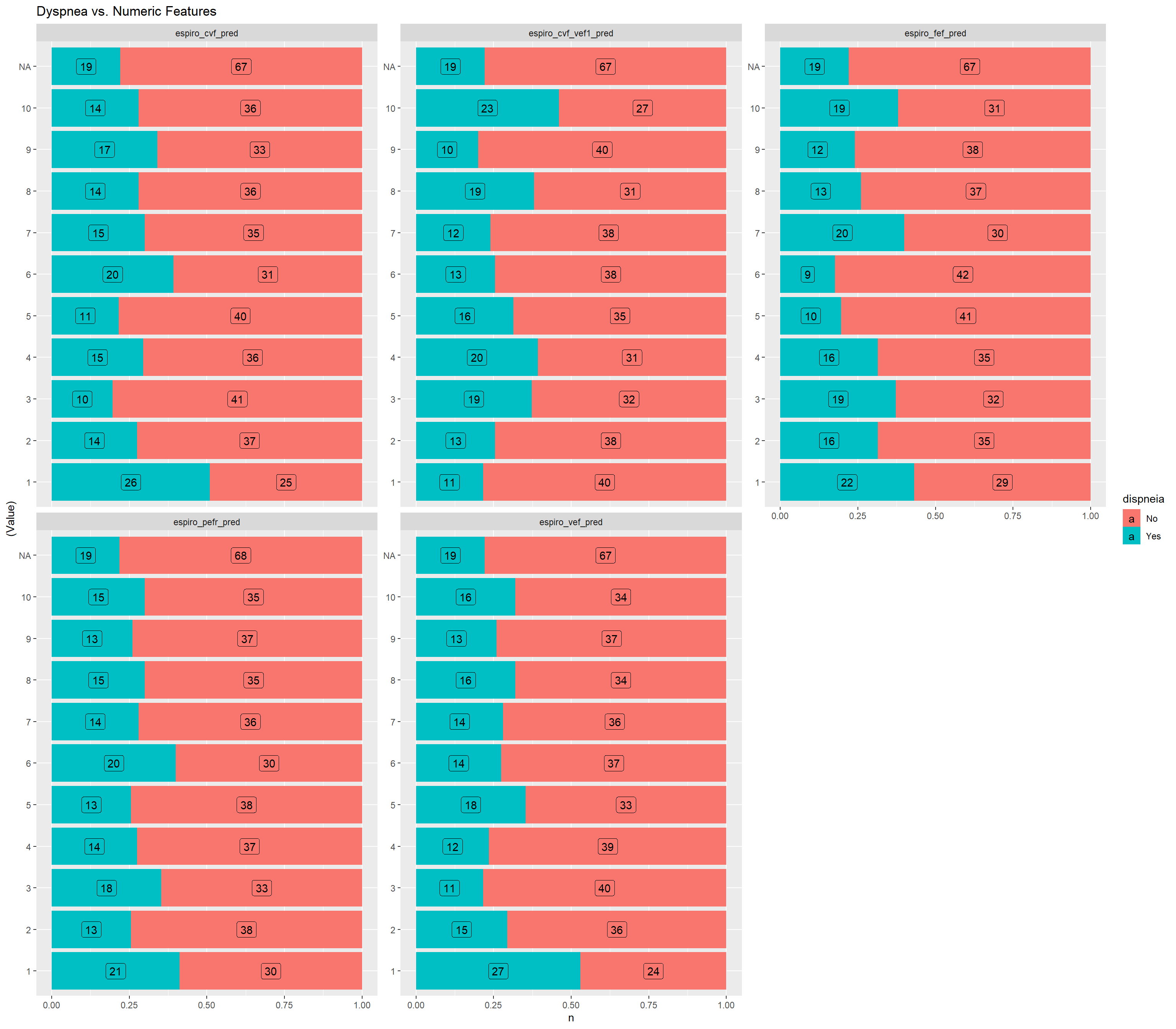
Checking the distribution of categorical features
contagens <-
dispneia_train |>
select(c(where(is.character),
dispneia)) |>
pivot_longer(-dispneia,
names_to = "Feature",
values_to = "Value") |>
count(dispneia,
Feature,
Value)
# tabela
contagens |>
pivot_wider(names_from = dispneia,
values_from = n) |>
DT::datatable()contagens |>
ggplot(aes(y = Value,
x = n,
fill = dispneia)) +
geom_col(position = "fill") +
geom_label(aes(label = n),
position = position_fill(vjust = 0.5)) +
facet_wrap(~Feature,
scales = "free_y",
ncol = 3) +
ggtitle("Dyspnea vs. Categorical Features")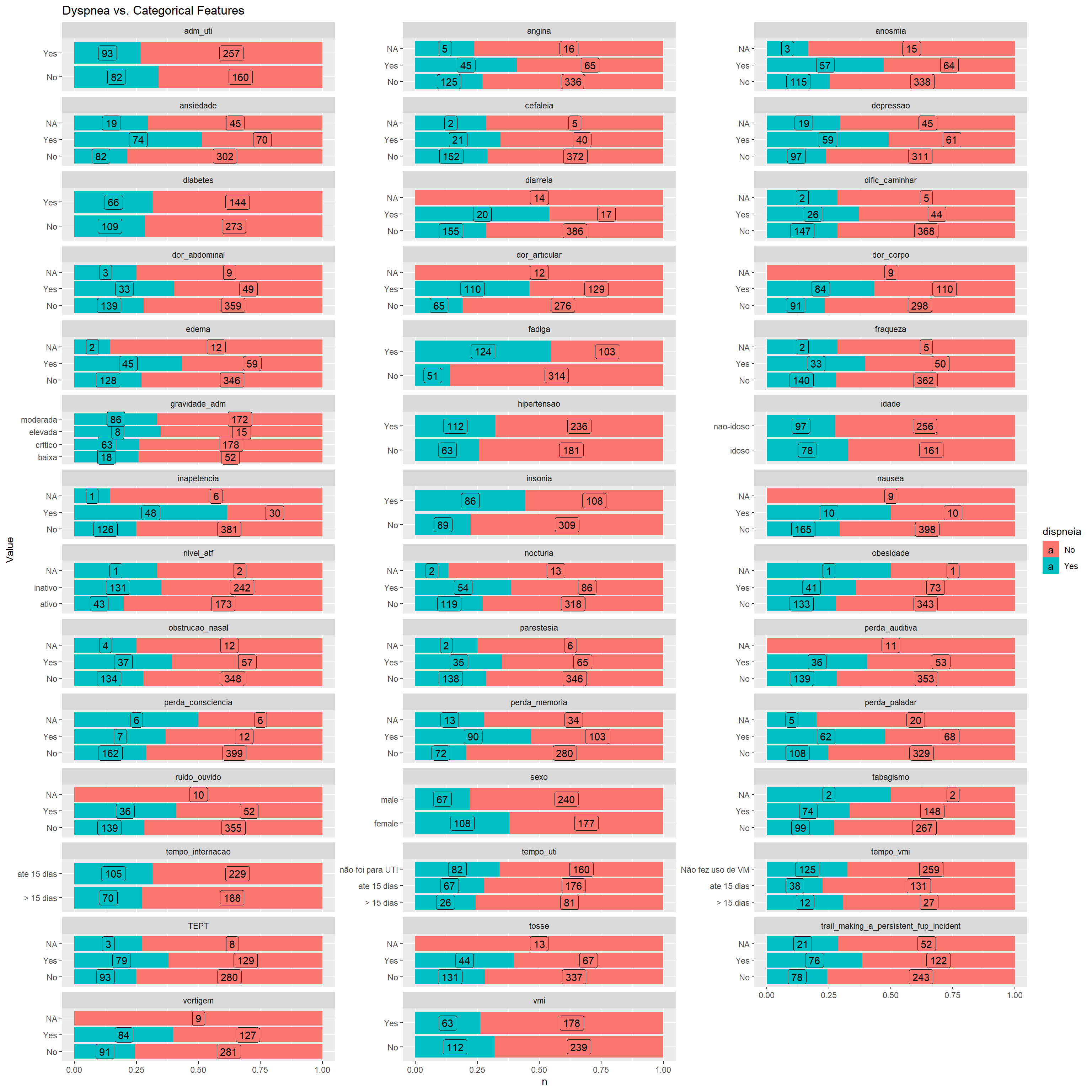
Preprocessing and Feature Engineering Steps using Recipes package
Recipe
For numeric features, the values were normalized, multicollineariry higher than 0.8 were removed and missing values were imputed with median.
For nominal features, missing values were imputed with more frequent value, and all features were transformed in a dummy feature.
Features containing only zero were removed.
The processed dataset can be seen below.
dispneia_recipe <-
recipe(formula = dispneia ~ ., dispneia_train) |>
step_normalize(all_numeric_predictors()) |> # normalize all continuous features
step_corr(all_numeric_predictors(), threshold = .8) |> # multicollinearity <- r > 0.8
step_impute_median(all_numeric_predictors()) |> # Processing missing continuous data - imputing median
step_impute_mode(all_nominal_predictors()) |> # Processing missing categorical data - imputing more frequent class
step_zv(all_predictors()) |> # removing variables that contain only zero (zero variance filter)
step_dummy(all_nominal_predictors()) # dummy - nominal predictors
# Checking the result of the recipe (and whether the steps worked!)
dispneia_recipe |>
prep() |>
bake(new_data = dispneia_train) |>
glimpse()## Rows: 592
## Columns: 50
## $ espiro_cvf_pred <dbl> 0.75971560, -0.93614868, -1…
## $ espiro_cvf_vef1_pred <dbl> 0.02173529, -0.18006391, 0.…
## $ espiro_pefr_pred <dbl> -0.5976043, -2.0833943, 0.2…
## $ espiro_fef_pred <dbl> 0.51947489, -1.17275315, -0…
## $ dispneia <fct> No, No, No, No, No, No, No,…
## $ idade_nao.idoso <dbl> 1, 0, 1, 0, 1, 0, 1, 0, 0, …
## $ sexo_male <dbl> 1, 0, 1, 1, 1, 1, 0, 1, 0, …
## $ adm_uti_Yes <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, …
## $ tempo_uti_ate.15.dias <dbl> 0, 1, 0, 1, 0, 0, 0, 1, 1, …
## $ tempo_uti_não.foi.para.UTI <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ vmi_Yes <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 0, …
## $ tempo_vmi_ate.15.dias <dbl> 0, 1, 0, 1, 0, 1, 1, 0, 0, …
## $ tempo_vmi_Não.fez.uso.de.VM <dbl> 0, 0, 0, 0, 0, 0, 0, 1, 1, …
## $ tosse_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ fraqueza_Yes <dbl> 1, 0, 1, 0, 0, 0, 0, 0, 0, …
## $ dific_caminhar_Yes <dbl> 0, 0, 1, 0, 0, 0, 0, 0, 0, …
## $ angina_Yes <dbl> 0, 0, 1, 0, 0, 0, 1, 0, 0, …
## $ dor_articular_Yes <dbl> 1, 1, 1, 0, 0, 0, 1, 0, 0, …
## $ dor_abdominal_Yes <dbl> 0, 0, 0, 1, 0, 0, 0, 0, 0, …
## $ diarreia_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ nausea_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ perda_consciencia_Yes <dbl> 0, 1, 0, 0, 0, 0, 0, 0, 0, …
## $ inapetencia_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ parestesia_Yes <dbl> 1, 0, 1, 0, 0, 1, 0, 0, 0, …
## $ obstrucao_nasal_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 1, 0, …
## $ perda_paladar_Yes <dbl> 1, 0, 0, 1, 0, 0, 0, 0, 0, …
## $ anosmia_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ fadiga_Yes <dbl> 0, 1, 0, 1, 1, 0, 1, 0, 0, …
## $ cefaleia_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ trail_making_a_persistent_fup_incident_Yes <dbl> 1, 0, 0, 0, 0, 0, 1, 0, 0, …
## $ perda_memoria_Yes <dbl> 0, 0, 0, 0, 0, 0, 1, 0, 1, …
## $ TEPT_Yes <dbl> 0, 0, 1, 1, 0, 0, 1, 0, 0, …
## $ perda_auditiva_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ ruido_ouvido_Yes <dbl> 0, 0, 1, 0, 0, 0, 1, 0, 0, …
## $ vertigem_Yes <dbl> 0, 1, 1, 1, 0, 0, 0, 0, 0, …
## $ edema_Yes <dbl> 1, 0, 0, 0, 0, 1, 1, 0, 0, …
## $ nocturia_Yes <dbl> 0, 0, 0, 0, 0, 1, 1, 1, 0, …
## $ dor_corpo_Yes <dbl> 0, 0, 0, 0, 1, 1, 1, 1, 1, …
## $ insonia_Yes <dbl> 0, 1, 0, 0, 0, 0, 1, 0, 1, …
## $ depressao_Yes <dbl> 0, 0, 0, 1, 0, 0, 0, 0, 0, …
## $ ansiedade_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ tempo_internacao_ate.15.dias <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 1, …
## $ tabagismo_Yes <dbl> 0, 0, 0, 0, 0, 0, 1, 0, 0, …
## $ obesidade_Yes <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ hipertensao_Yes <dbl> 1, 1, 0, 1, 1, 1, 1, 0, 1, …
## $ diabetes_Yes <dbl> 1, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ gravidade_adm_critico <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 0, …
## $ gravidade_adm_elevada <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, …
## $ gravidade_adm_moderada <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 1, …
## $ nivel_atf_inativo <dbl> 0, 1, 1, 1, 0, 1, 0, 1, 1, …Definition of cross-validation
dispneia_resamples <-
vfold_cv(dispneia_train,
v = 5,
strata = dispneia) # 5 folds
dispneia_resamples$splits## [[1]]
## <Analysis/Assess/Total>
## <473/119/592>
##
## [[2]]
## <Analysis/Assess/Total>
## <473/119/592>
##
## [[3]]
## <Analysis/Assess/Total>
## <474/118/592>
##
## [[4]]
## <Analysis/Assess/Total>
## <474/118/592>
##
## [[5]]
## <Analysis/Assess/Total>
## <474/118/592>Modeling
XGBoost
The hyperameters tuned were:
- boost_tree;
- min_n: An integer for the minimum number of data points in a node that is required for the node to be split further;
- mtry: A number for the number (or proportion) of predictors that will be randomly sampled at each split when creating the tree models (specific engines only);
- trees: An integer for the number of trees contained in the ensemble;
- tree_depth: An integer for the maximum depth of the tree (i.e. number of splits) (specific engines only);
- learn_rate: A number for the rate at which the boosting algorithm adapts from iteration-to-iteration (specific engines only). This is sometimes referred to as the shrinkage parameter;
- loss_reduction: A number for the reduction in the loss function required to split further (specific engines only);
- sample_size: A number for the number (or proportion) of data that is exposed to the fitting routine. For xgboost, the sampling is done at each iteration while C5.0 samples once during training.
The tuning process were done in 5 steps.
Step 1 - Number of trees (trees) and learn rate (learn_rate)
Model
# Modelo
dispneia_xgb_model <-
boost_tree(
min_n = 15,
mtry = 0.8,
trees = tune(),
tree_depth = 5,
learn_rate = tune(),
loss_reduction = 0,
sample_size = 0.8
) |>
set_mode("classification") |>
set_engine("xgboost", count = FALSE)Workflow
# Workflow
dispneia_xgb_wf <-
workflow() |>
add_model(dispneia_xgb_model) |>
add_recipe(dispneia_recipe)Tuning
# Tune
dispneia_grid_xgb <-
expand.grid(
trees = seq(25, 800, 25),
learn_rate = seq(0.01, 0.4,0.1)
)
dispneia_grid_xgb |>
DT::datatable()doParallel::registerDoParallel(4)
dispneia_xgb_tune_grid <-
tune_grid(
dispneia_xgb_wf,
resamples = dispneia_resamples,
grid = dispneia_grid_xgb,
metrics = metric_set(roc_auc, accuracy, sensitivity, specificity),
verbose = TRUE
)autoplot(dispneia_xgb_tune_grid) +
theme_bw() +
theme(legend.position = "top")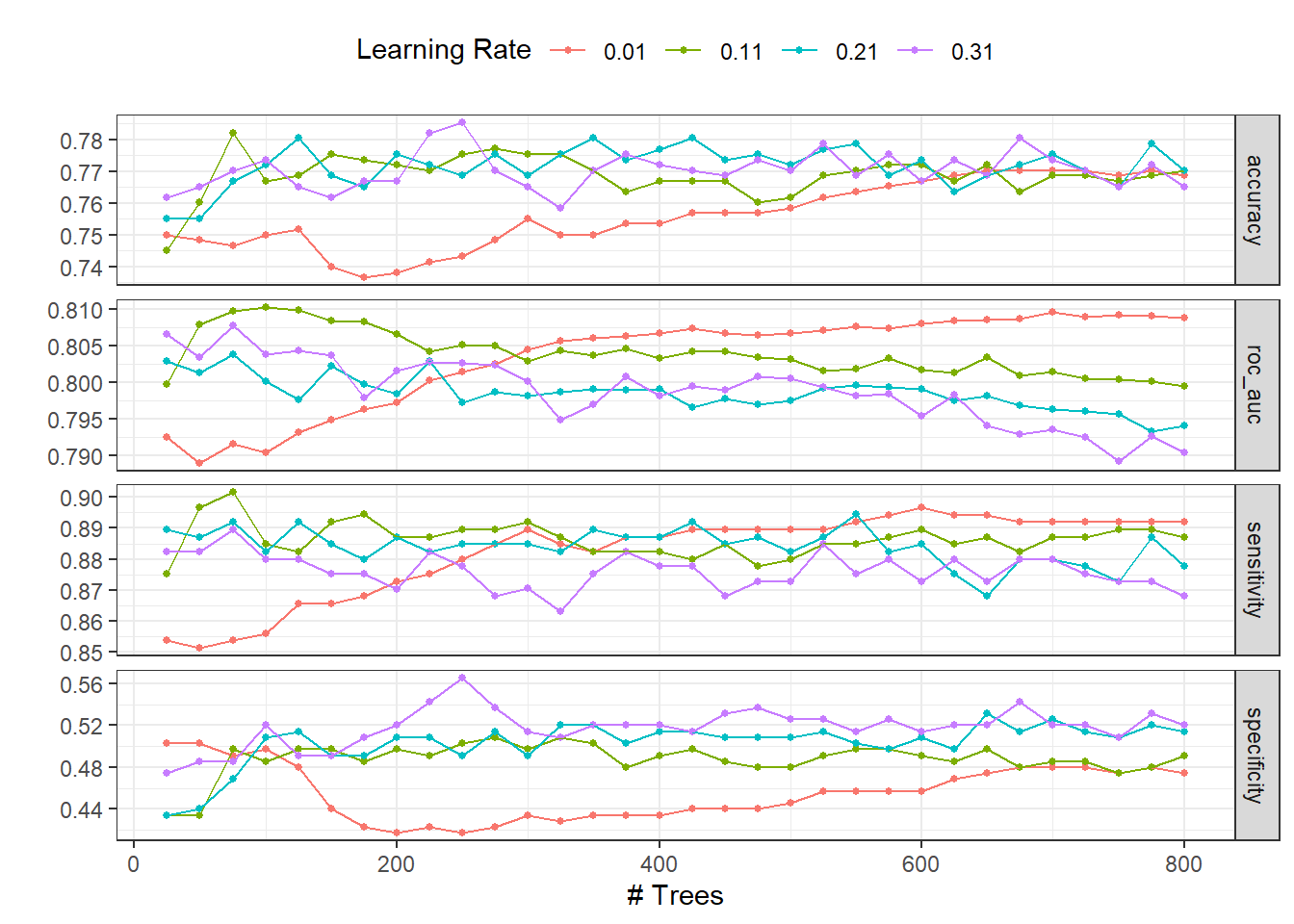
collect_metrics(dispneia_xgb_tune_grid) |>
filter(.metric == "roc_auc") |>
filter(mean == max(mean))## # A tibble: 1 × 8
## trees learn_rate .metric .estimator mean n std_err .config
## <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 100 0.11 roc_auc binary 0.810 5 0.0163 Preprocessor1_Model036dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")## # A tibble: 1 × 3
## trees learn_rate .config
## <dbl> <dbl> <chr>
## 1 100 0.11 Preprocessor1_Model036show_best(dispneia_xgb_tune_grid, n = 6)## # A tibble: 6 × 8
## trees learn_rate .metric .estimator mean n std_err .config
## <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 100 0.11 roc_auc binary 0.810 5 0.0163 Preprocessor1_Model036
## 2 125 0.11 roc_auc binary 0.810 5 0.0161 Preprocessor1_Model037
## 3 75 0.11 roc_auc binary 0.810 5 0.0163 Preprocessor1_Model035
## 4 700 0.01 roc_auc binary 0.810 5 0.0166 Preprocessor1_Model028
## 5 750 0.01 roc_auc binary 0.809 5 0.0162 Preprocessor1_Model030
## 6 775 0.01 roc_auc binary 0.809 5 0.0164 Preprocessor1_Model031dispneia_best_step1 <-
dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")# best hiperparameters
glue::glue("The best number of trees was {dispneia_best_step1$trees} and learn_rate was {dispneia_best_step1$learn_rate}.")## The best number of trees was 100 and learn_rate was 0.11.Step 2 - Minimal node size (min_n) and depth tree (tree_depth)
Model
# Modelo
dispneia_xgb_model <-
boost_tree(
min_n = tune(),
mtry = 0.8,
trees = 25,
tree_depth = tune(),
learn_rate = 0.31,
loss_reduction = 0,
sample_size = 0.8
) |>
set_mode("classification") |>
set_engine("xgboost", count = FALSE)Workflow
# Workflow
dispneia_xgb_wf <-
workflow() |>
add_model(dispneia_xgb_model) |>
add_recipe(dispneia_recipe)Tuning
# Tune - min_n and tree_depth
dispneia_grid_xgb <-
expand.grid(
min_n = seq(1,16,2),
tree_depth = seq(1,17,4)
)
dispneia_grid_xgb |>
DT::datatable()doParallel::registerDoParallel(4)
dispneia_xgb_tune_grid <-
tune_grid(
dispneia_xgb_wf,
resamples = dispneia_resamples,
grid = dispneia_grid_xgb,
metrics = metric_set(roc_auc, accuracy, sensitivity, specificity),
verbose = TRUE
)autoplot(dispneia_xgb_tune_grid) +
theme_bw() +
theme(legend.position = "top")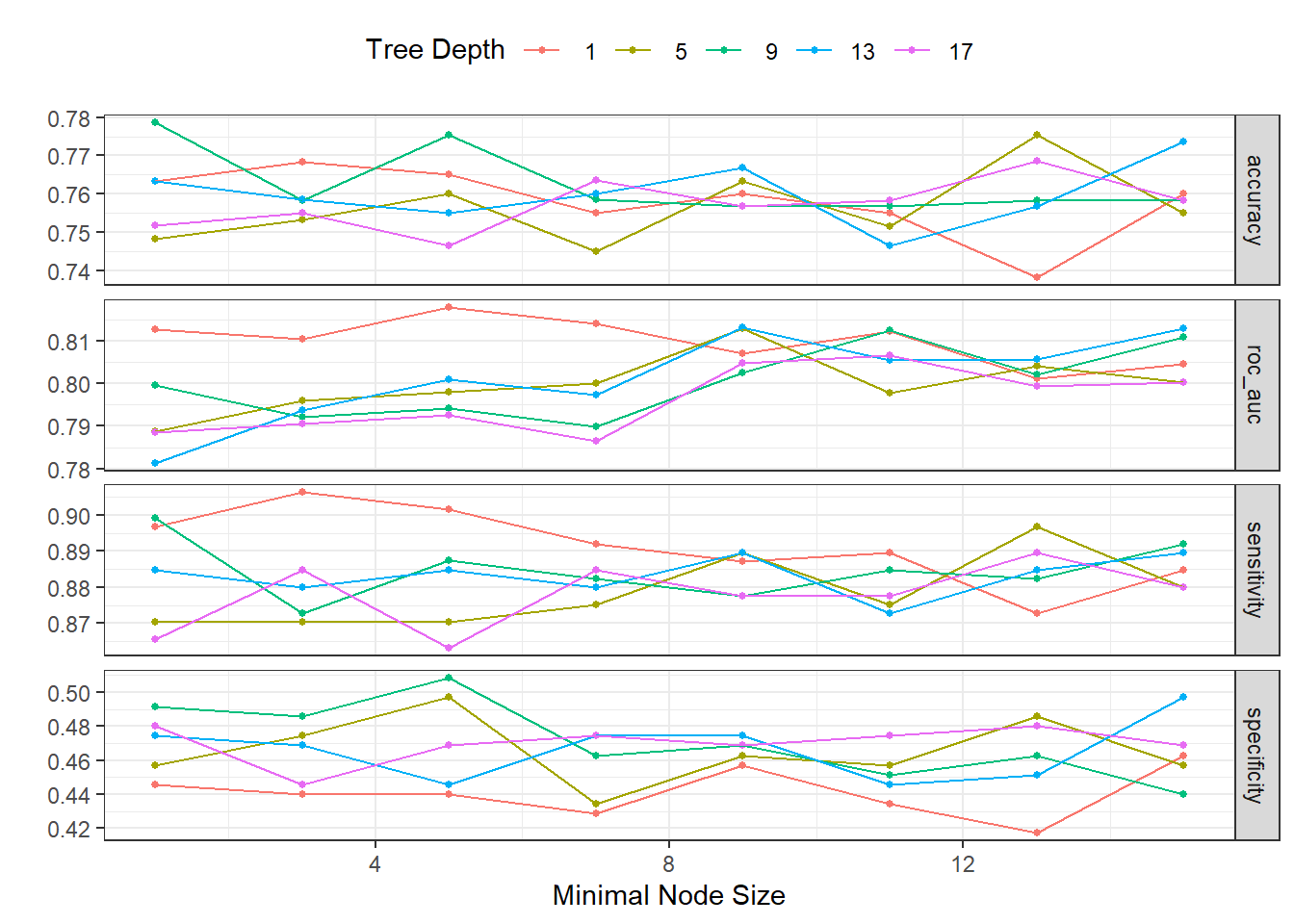
collect_metrics(dispneia_xgb_tune_grid) |>
filter(.metric == "roc_auc") |>
filter(mean == max(mean))## # A tibble: 1 × 8
## min_n tree_depth .metric .estimator mean n std_err .config
## <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 5 1 roc_auc binary 0.818 5 0.0209 Preprocessor1_Model03dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")## # A tibble: 1 × 3
## min_n tree_depth .config
## <dbl> <dbl> <chr>
## 1 5 1 Preprocessor1_Model03show_best(dispneia_xgb_tune_grid, n = 6)## # A tibble: 6 × 8
## min_n tree_depth .metric .estimator mean n std_err .config
## <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 5 1 roc_auc binary 0.818 5 0.0209 Preprocessor1_Model03
## 2 7 1 roc_auc binary 0.814 5 0.0213 Preprocessor1_Model04
## 3 9 13 roc_auc binary 0.813 5 0.0161 Preprocessor1_Model29
## 4 9 5 roc_auc binary 0.813 5 0.0148 Preprocessor1_Model13
## 5 15 13 roc_auc binary 0.813 5 0.0153 Preprocessor1_Model32
## 6 1 1 roc_auc binary 0.813 5 0.0213 Preprocessor1_Model01dispneia_best_step2 <-
dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")# best hiperparameters
glue::glue("The best minimal node size was {dispneia_best_step2$min_n} and depth tree was {dispneia_best_step2$tree_depth}.")## The best minimal node size was 5 and depth tree was 1.Step 3 - Number for the reduction in the loss function required to split further (loss_reduction)
Model
# Modelo
dispneia_xgb_model <-
boost_tree(
min_n = 11,
mtry = 0.8,
trees = 25,
tree_depth = 1,
learn_rate = 0.31,
loss_reduction = tune(),
sample_size = 0.8
) |>
set_mode("classification") |>
set_engine("xgboost", count = FALSE)Workflow
# Workflow
dispneia_xgb_wf <-
workflow() |>
add_model(dispneia_xgb_model) |>
add_recipe(dispneia_recipe)Tuning
# Tune - min_n and tree_depth
dispneia_grid_xgb <-
expand.grid(
loss_reduction = seq(1, 4, 0.04)
)
dispneia_grid_xgb |>
DT::datatable()doParallel::registerDoParallel(4)
dispneia_xgb_tune_grid <-
tune_grid(
dispneia_xgb_wf,
resamples = dispneia_resamples,
grid = dispneia_grid_xgb,
metrics = metric_set(roc_auc, accuracy, sensitivity, specificity),
verbose = TRUE
)autoplot(dispneia_xgb_tune_grid) +
theme_bw() 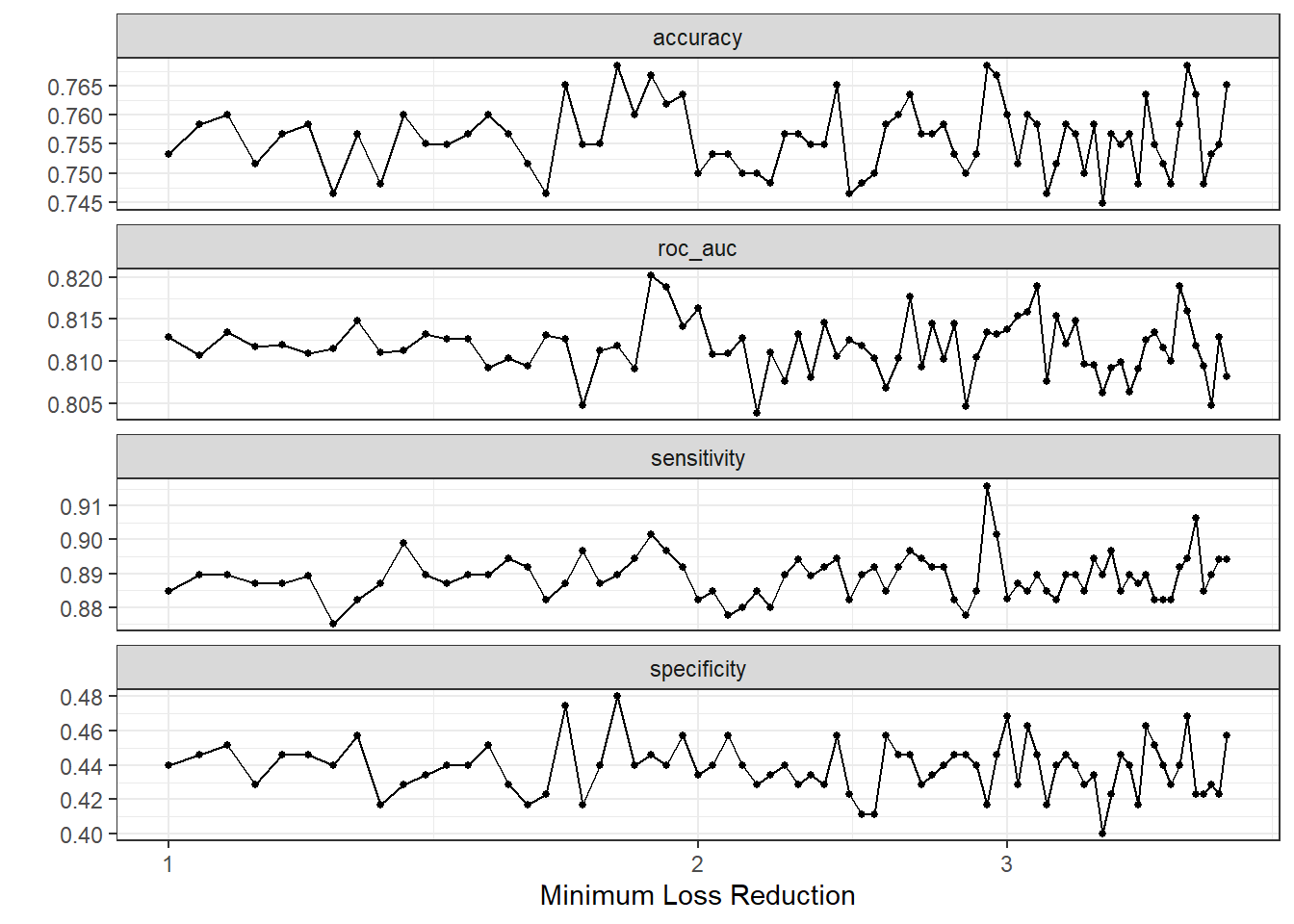
collect_metrics(dispneia_xgb_tune_grid) |>
filter(.metric == "roc_auc") |>
filter(mean == max(mean))## # A tibble: 1 × 7
## loss_reduction .metric .estimator mean n std_err .config
## <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 1.88 roc_auc binary 0.820 5 0.0195 Preprocessor1_Model23dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")## # A tibble: 1 × 2
## loss_reduction .config
## <dbl> <chr>
## 1 1.88 Preprocessor1_Model23show_best(dispneia_xgb_tune_grid, n = 6)## Warning: No value of `metric` was given; metric 'roc_auc' will be used.## # A tibble: 6 × 7
## loss_reduction .metric .estimator mean n std_err .config
## <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 1.88 roc_auc binary 0.820 5 0.0195 Preprocessor1_Model23
## 2 3.76 roc_auc binary 0.819 5 0.0168 Preprocessor1_Model70
## 3 3.12 roc_auc binary 0.819 5 0.0207 Preprocessor1_Model54
## 4 1.92 roc_auc binary 0.819 5 0.0206 Preprocessor1_Model24
## 5 2.64 roc_auc binary 0.818 5 0.0205 Preprocessor1_Model42
## 6 2 roc_auc binary 0.816 5 0.0205 Preprocessor1_Model26dispneia_best_step3 <-
dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")# best hiperparameters
glue::glue("The best loss_reduction was {dispneia_best_step3$loss_reduction}.")## The best loss_reduction was 1.88.Step 4 - Number (or proportion) of predictors that will be randomly sampled (mtry) and number (or proportion) of data that is exposed to the fitting routine (sample_size)
Model
# Modelo
dispneia_xgb_model <-
boost_tree(
min_n = 11,
mtry = tune(),
trees = 25,
tree_depth = 1,
learn_rate = 0.31,
loss_reduction = 1.36,
sample_size = tune()
) |>
set_mode("classification") |>
set_engine("xgboost", count = FALSE)Workflow
# Workflow
dispneia_xgb_wf <-
workflow() |>
add_model(dispneia_xgb_model) |>
add_recipe(dispneia_recipe)Tuning
# Tune - min_n and tree_depth
dispneia_grid_xgb <-
expand.grid(
mtry = seq(0.2, 1.0, 0.2),
sample_size = seq(0.2, 1.0,by = 0.2)
)
dispneia_grid_xgb |>
DT::datatable()doParallel::registerDoParallel(4)
dispneia_xgb_tune_grid <-
tune_grid(
dispneia_xgb_wf,
resamples = dispneia_resamples,
grid = dispneia_grid_xgb,
metrics = metric_set(roc_auc, accuracy, sensitivity, specificity),
verbose = TRUE
)autoplot(dispneia_xgb_tune_grid) +
theme_bw() +
theme(legend.position = "top")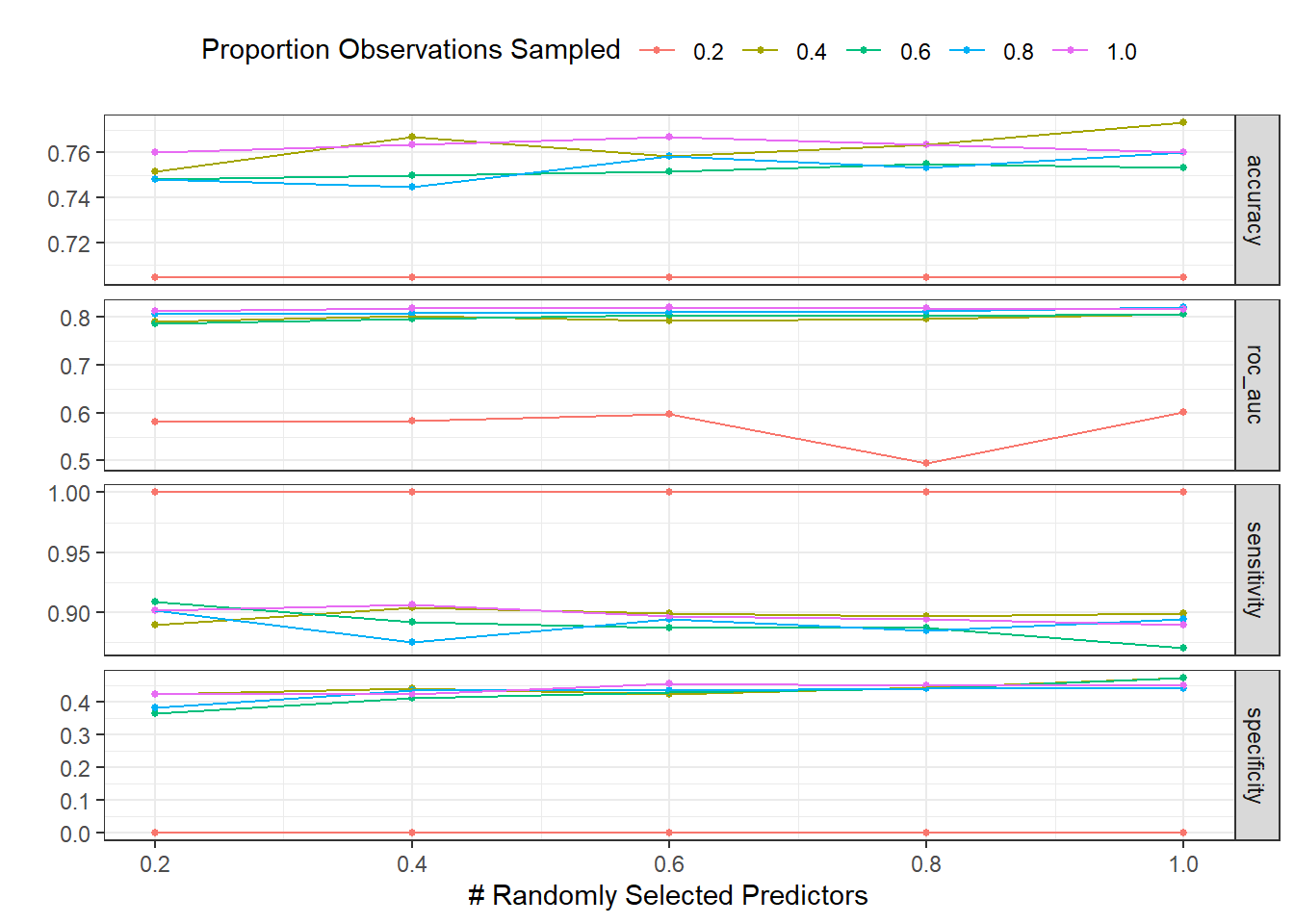
collect_metrics(dispneia_xgb_tune_grid) |>
filter(.metric == "roc_auc") |>
filter(mean == max(mean))## # A tibble: 1 × 8
## mtry sample_size .metric .estimator mean n std_err .config
## <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 0.6 1 roc_auc binary 0.820 5 0.0217 Preprocessor1_Model23dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")## # A tibble: 1 × 3
## mtry sample_size .config
## <dbl> <dbl> <chr>
## 1 0.6 1 Preprocessor1_Model23show_best(dispneia_xgb_tune_grid, n = 6)## Warning: No value of `metric` was given; metric 'roc_auc' will be used.## # A tibble: 6 × 8
## mtry sample_size .metric .estimator mean n std_err .config
## <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 0.6 1 roc_auc binary 0.820 5 0.0217 Preprocessor1_Model23
## 2 1 0.8 roc_auc binary 0.819 5 0.0197 Preprocessor1_Model20
## 3 0.4 1 roc_auc binary 0.818 5 0.0218 Preprocessor1_Model22
## 4 0.8 1 roc_auc binary 0.818 5 0.0215 Preprocessor1_Model24
## 5 1 1 roc_auc binary 0.817 5 0.0225 Preprocessor1_Model25
## 6 0.2 1 roc_auc binary 0.812 5 0.0227 Preprocessor1_Model21dispneia_best_step4 <-
dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")# best hiperparameters
glue::glue("The best number (or proportion) of predictors that will be randomly sampled (mtry) was {dispneia_best_step4$mtry} and the number (or proportion) of data that is exposed to the fitting routine (sample size) was {dispneia_best_step4$sample_size}.")## The best number (or proportion) of predictors that will be randomly sampled (mtry) was 0.6 and the number (or proportion) of data that is exposed to the fitting routine (sample size) was 1.Step 5 - Training again learing rate and trees with lower values
Model
# Modelo
dispneia_xgb_model <-
boost_tree(
min_n = 11,
mtry = 0.8,
trees = tune(),
tree_depth = 1,
learn_rate = tune(),
loss_reduction = 1.36,
sample_size = 0.8
) |>
set_mode("classification") |>
set_engine("xgboost", count = FALSE)Workflow
# Workflow
dispneia_xgb_wf <-
workflow() |>
add_model(dispneia_xgb_model) |>
add_recipe(dispneia_recipe)Tuning
# Tune - min_n and tree_depth
dispneia_grid_xgb <-
expand.grid(
trees = seq(15, 700, 25),
learn_rate = seq(0.02, 0.1, 0.02)
)
dispneia_grid_xgb |>
DT::datatable()doParallel::registerDoParallel(4)
dispneia_xgb_tune_grid <-
tune_grid(
dispneia_xgb_wf,
resamples = dispneia_resamples,
grid = dispneia_grid_xgb,
metrics = metric_set(roc_auc, accuracy, sensitivity, specificity),
verbose = TRUE
)autoplot(dispneia_xgb_tune_grid) +
theme_bw() +
theme(legend.position = "top")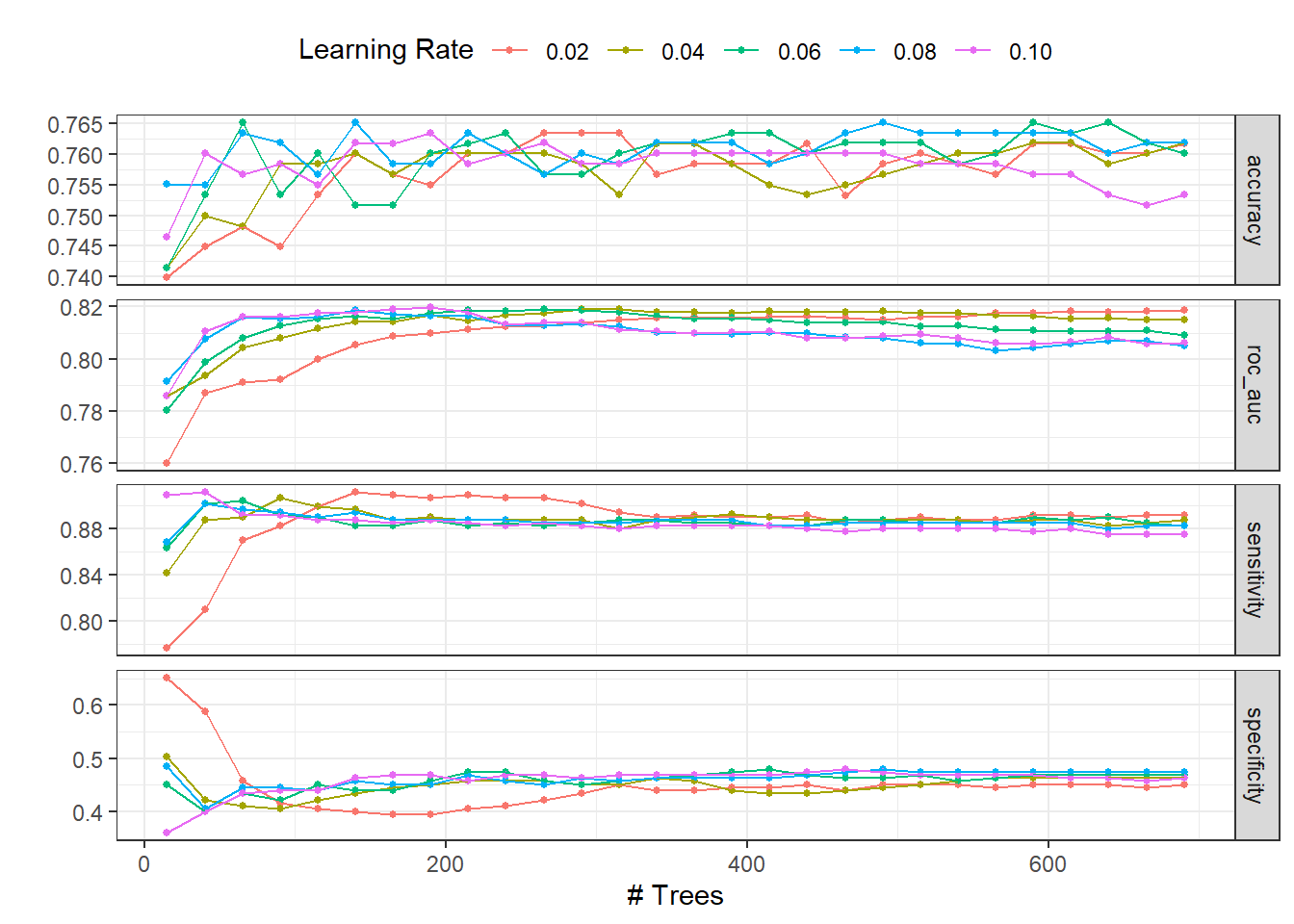
collect_metrics(dispneia_xgb_tune_grid) |>
filter(.metric == "roc_auc") |>
filter(mean == max(mean))## # A tibble: 1 × 8
## trees learn_rate .metric .estimator mean n std_err .config
## <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 190 0.1 roc_auc binary 0.820 5 0.0169 Preprocessor1_Model120dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")## # A tibble: 1 × 3
## trees learn_rate .config
## <dbl> <dbl> <chr>
## 1 190 0.1 Preprocessor1_Model120show_best(dispneia_xgb_tune_grid, n = 6)## Warning: No value of `metric` was given; metric 'roc_auc' will be used.## # A tibble: 6 × 8
## trees learn_rate .metric .estimator mean n std_err .config
## <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
## 1 190 0.1 roc_auc binary 0.820 5 0.0169 Preprocessor1_Model120
## 2 165 0.1 roc_auc binary 0.819 5 0.0175 Preprocessor1_Model119
## 3 290 0.04 roc_auc binary 0.819 5 0.0195 Preprocessor1_Model040
## 4 265 0.06 roc_auc binary 0.819 5 0.0191 Preprocessor1_Model067
## 5 315 0.04 roc_auc binary 0.819 5 0.0196 Preprocessor1_Model041
## 6 290 0.06 roc_auc binary 0.819 5 0.0187 Preprocessor1_Model068dispneia_best_step5 <-
dispneia_xgb_tune_grid |>
select_best(metric = "roc_auc")# best hiperparameters
glue::glue("Following tunning again, the best number of trees was {dispneia_best_step5$trees} and learn_rate was {dispneia_best_step5$learn_rate}.")## Following tunning again, the best number of trees was 190 and learn_rate was 0.1.Final model performance
Model
# Model
dispneia_xgb_final_model <-
boost_tree(
min_n = 11,
mtry = 0.8,
trees = 565,
tree_depth = 1,
learn_rate = 0.02,
loss_reduction = 1.36,
sample_size = 0.8
) |>
set_mode("classification") |>
set_engine("xgboost", count = FALSE)Workflow
# Workflow
dispneia_xgb_final_wf <-
workflow() |>
add_model(dispneia_xgb_final_model) |>
add_recipe(dispneia_recipe)Tuning
# Tune - min_n and tree_depth
dispneia_best_params_xgb <-
expand.grid(
min_n = 11,
mtry = 0.8,
trees = 565,
tree_depth = 1,
learn_rate = 0.02,
loss_reduction = 1.36,
sample_size = 0.8
)
dispneia_best_params_xgb |>
DT::datatable()# Desempenho do modelo finaL ----------------------------------------------
dispneia_xgb_final_wf <-
dispneia_xgb_final_wf |>
finalize_workflow(dispneia_best_params_xgb)
dispneia_xgb_last_fit <-
last_fit(dispneia_xgb_final_wf, dispneia_initial_split)
collect_predictions(dispneia_xgb_last_fit) |>
mutate(modelo = "XGBoost") |>
group_by(modelo) |>
roc_curve(dispneia, .pred_No) |>
autoplot() +
theme_bw() +
theme(legend.position = "none")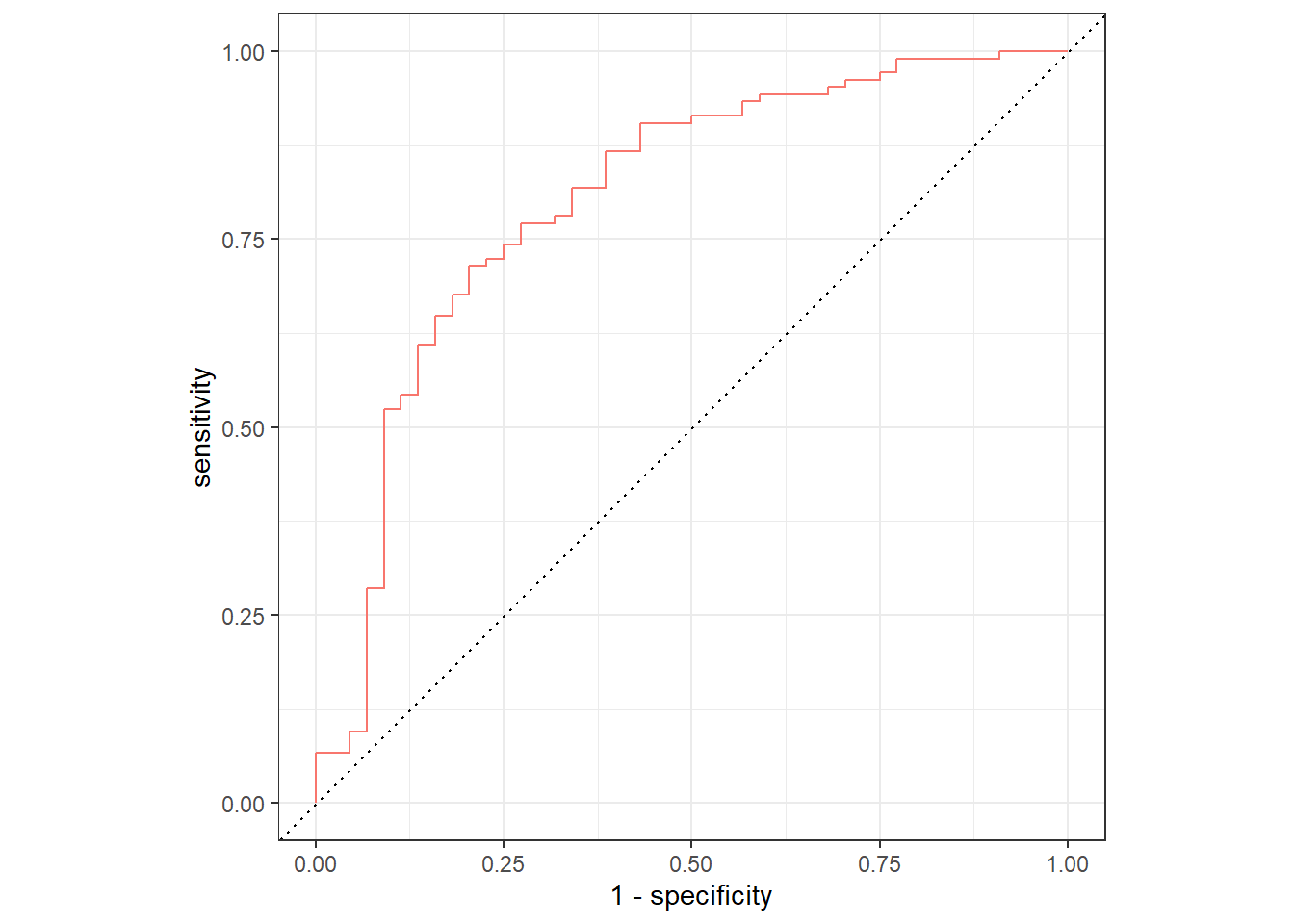
# best hiperparameters
best_roc <- show_best(dispneia_xgb_last_fit)
glue::glue("The best ROC was {round(best_roc$mean,2)}!")## The best ROC was 0.8!dispneia_xgb_last_fit_model <- dispneia_xgb_last_fit$.workflow[[1]]$fit$fit
dispneia_xgb_vip <-
extract_fit_engine(dispneia_xgb_last_fit)
vip::vip(dispneia_xgb_vip)+
theme_bw()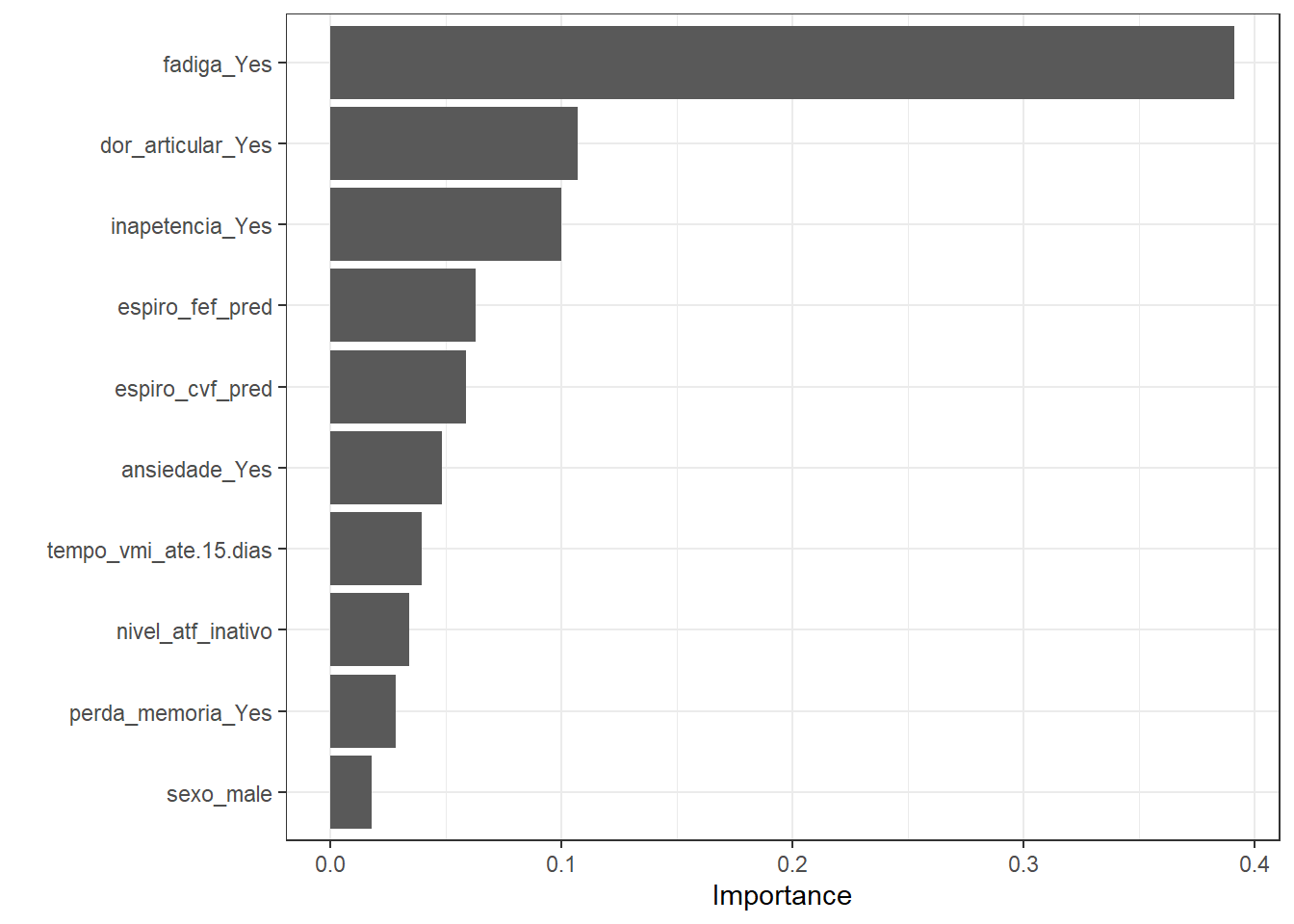
Conclusions
Finally, XGBoost was effective to predict dyspnea in COVID-19 survivors 6-9 months after hospital discharge. Thus, XGBoost implementation in a clinical setting might aid health professionals in the early management of COVID-19 survivors at risk of future dyspnea.
It is noteworthy that the analyses performed herein are not intended to be used in practice. However, these analysis reinforce the utilization of ML algorithm in the health care.
Hope you enjoyed it!😜!
